Everyone likes free biological sequence search.
A free IP sequence search usually involves the following steps: search the Genbank patent divisions on the NCBI BLAST web site, go through the alignments one by one, and lookup related patent information on the web.
Bad idea. Here are four reasons why.
1. Free sources are incomplete.
Why waste your time searching at all if you aren’t searching the complete register? The Genbank December 2014 release (205) had just under 180 million sequences. The GQ-Pat database had over 280 million sequences at that time, and can be searched right alongside the 180 million sequences in Genbank.
2. Free searches are stuck in the wrong algorithms.
The most common error in IP sequence searching is using a publicly available algorithm like BLAST to determine legal relevance. It’s important to understand that BLAST, the most popular sequence search algorithm, has been created with biology in mind. It answers questions like: “is there an equivalent sequence in another species?” It is the wrong algorithm to answer an IP-related questions such as, “find all sequences in the database that are 70% or more identical to my query sequence.” For that, an algorithm like GenePAST is more appropriate.
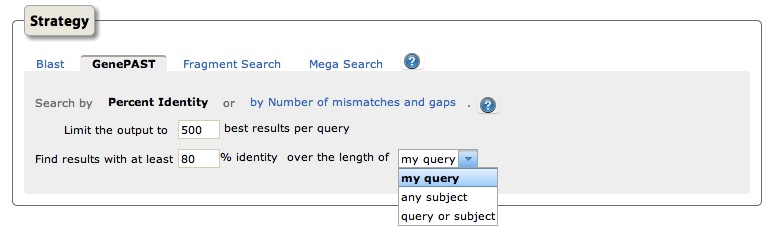
3. Free searches take too much effort.
Public services present the outcome of a search as a long, static, list of alignments. From that list there is no easy way to filter the relevant hits and retrieve information about the related IP documents. A common solution is to print out everything, go through the hits one by one, and look up related patent information on the web. Findings are scribbled on to the printout or put into a spreadsheet. This approach is labor intensive, error-prone, and very inflexible. When the question changes the entire procedure has to be repeated from scratch.
GenomeQuest presents the outcome of a search as an interactive and fully queryable list that contains information about the alignments, the sequences, and the related IP documents. With a couple of clicks, users can find all hits with 70% identity or more over at least 500 nucleotides, where the sequence is claimed in a granted patent with the filing date before January 2010.
4. Free searches are not private.
Anyone can see the communication between your web browser and an unencrypted server like NCBI’s. And here you are running searches of the most confidential nature on the open Internet! Use a trusted, HIPAA-complaint, SAS II compliant service to ensure the security of your inventions.

